How to Split Reads Into Forward and Reverse
nine.i - The FastQ file format
Results of Sanger sequencing are unremarkably fasta files (obtained from processing chromatograms). Most high-throughput sequencing machines output fastq files, the "de facto" current standard in HTS. Like fasta, fastq files are simply text files, but where each cake of information (a sequenced DNA fragment, or read) in this format is encoded equally 4 lines:
@read_identifier read_sequence + separator line base_qualities For example, here you accept 8 lines of a fastq file, corresponding to ii sequences:
@HWI-M01876:76:000000000-AF16W:ane:1101:10853:k i:Northward:0:CGTGACAGAT NTGTACTTCATCCGAAACTCGTGCTCATCTCTGCTCAGATCGGAAGAGCACACGTCTGAACTCCAGTCACCGTGAT + #8ABCFGGGFCEDCFGGGGGGGFFCGEFGGGGGGFGGGGGGGGDEFGGGGGGGGGGGGGGGGGFFFEGGGGGGGGF @HWI-M01876:76:000000000-AF16W:1:1101:16471:1000 1:N:0:CGTGAACTTG NTTCCAGATATTCGATGCATGTGCCGCTCCTGTCGGAGATCGGAAGAGCACACGTCTGAACTCCAGTCACCGTGAT + #8BCCGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGEGGGGFGGGGGGGGGGGGGGGGGGGGGGGGGG Each base has a quality character associated with it, representing how confidently the car identified (called) the base. The probability of error per base is given as a Phred score, calculated from an integer value (Q) derived from the quality character associated to the base. The probability of mistake is given by the Phred score using P(Q)=10^(-Q/ten). Useful reference values of Q include:
- Q=10 - xc% accuracy (0.1 mistake)
- Q=20 - 99% accurateness (0.01 mistake)
- Q=thirty - 99.9% accurateness (0.001 error)
- Q=xl - 99.99% accuracy (0.0001 error)
Although there'south theoretically no limit, Q unremarkably goes upwardly to around 40 in contempo illumina machines.
To obtain this Q value from the character associated to the quality of the base of operations, we take to know that each character (such as '#') has an ASCII decimal value associated (for example, '#' has a value of 35). The Q value of a character is the decimal value corresponding to the entry of that character in the ASCII table, subtracted past 33. For case Q('#') = 35 – 33.
Annotation: To sympathize why nosotros need to subtract 33, nosotros have to wait into the ASCII table below. We can see that the first visible character ('!') has decimal value 33. This allows visual inspection of qualities.
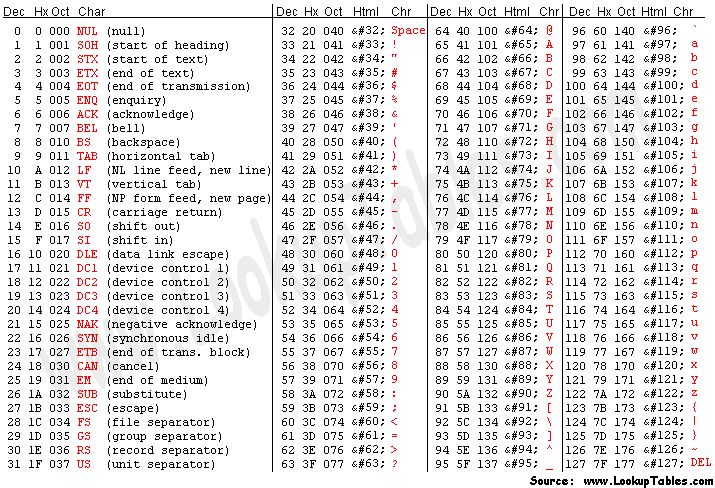
Looking at the get-go read of our fastq example, we can meet information technology starts with 'N' (unknown), with an associated quality graphic symbol '#'. To know how confident the machine was in reading that base of operations, we calculate:
- Q = 35 (ASCII decimal value of '#') - 33 (ASCII decimal value of '!') = ii
- p(two) = 10^(-2/10) = 63% (probability of error)
Given this probability of error, it is non surprising that the machine could non confidently say which base was in that position and therefore placed an 'N' in that position. It is adequately mutual that in the first bases the machine is withal calibrating, and sometimes there is less conviction in the chosen base.
TASK: Summate the probability of fault of the bases of the following read in the fastq format:
@SRR022885.1 BI:080102_SL-XAR_0001_FC201E9AAXX:6:one:752:593/1 CGTACCAATTATTCAACGTCGCCAGTTGCTTCATGT + IIIIIIIIII>IIIIIII@IIII.I+I>35I0I&+/ Note: Phred+33 (Sanger fastq) is the electric current standard format. Notwithstanding, with older illumina data (before 2009) preferred to start at the graphic symbol '@' (ASCII: 64) instead of '!'. This Phred+64 format is the old illumina fastq. Some tools (similar FastQC) can infer the format, while in others y'all need to specify.
QUESTION: What is the probability of fault of the outset base of the read?
Click Here to see the answer
The base of operations quality grapheme is 'I', which corresponds to the decimal 73 in the ASCII table.
Q = 73-33 = 40. P(twoscore) = ten^(-40/ten) = 10^-4 = 0.01% error.
QUESTION: What is the probability of error of the final base of the read?
Click Hither to encounter the answer
The base of operations quality character is '/', which corresponds to the decimal 47 in the ASCII table.
Q = 47-33 = fourteen. P(14) = ten^(-14/10) = x^-one.iv ~= 4% error.
QUESTION: If all bases of a ficticious automobile had a Q=xx (ane% probability of error), what would be the probability that 1 100bp read from that machine would be completely correct?
Click Here to run into the reply
P(correct)=(0.99)^100 ~= 36.six%!
This serves to exemplify that about reads in electric current sequencing machines are probable to have at least one base of operations incorrect.
Many sequencing machines tin can read both ends of a fragment. In this example, the car volition generate two paired fastq files, one with the forward reads and another with the opposite reads. Yous can find an example of this is the instance fastq files paired_end_example_1 (containing the forward reads) and paired_end_example_2 (containing the opposite reads). These fastq are paired considering the reads for the same fragment are in the aforementioned gild in the two files. For example, the first read in the forward fastq corresponds to the forward reading of the same fragment every bit the commencement read in the reverse fastq.
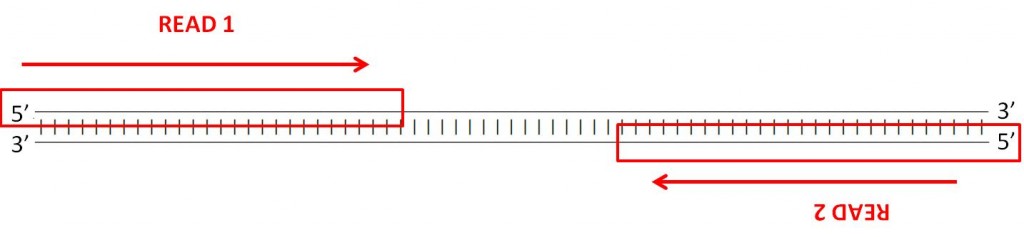
QUESTION: Inside the binder fastq_examples you can encounter several compressed fastq files. Uncompress the fastq files paired_end_example_1.fastq.gz and paired_end_example_2.fastq.gz that are in the folder fastq_examples (either past clicking on the files, or using gunzip on the command line). Open the uncompressed fastq files using any text editor (eg. kate). Tin y'all see a relationship betwixt the reads in both files?
Click Here to encounter the answer
The read identifiers are the same, in the aforementioned guild (though the sequences are not). This is because they are readings of the same fragment, one (_1) in the forward and another (_2) in the reverse direction. Often the indication of forrad and reverse is in the identifier itself.
NOTE: Assess how well yous achieved the learning outcome. For this, run into how well you lot responded to the unlike questions during the activities and also make the following questions to yourself.
-
How well practise you understand the content of a fastQ file?
-
Tin y'all manually calculate the probability of error associated to whatsoever given base in a fastq file?
-
Did y'all understand the divergence betwixt single-cease and paired-terminate reads?
9.ii - Quality Check of FastQ data
High Throughput Sequencing machines read thousands or millions of sequences in parallel. As y'all can imagine, this usually generates large fastq files, with millions of lines. Manually inspecting the quality of each read is out of the question. Specialized software has been developed to provide quality measures for fastq files generated by HTS machines. FastQC is a popular programme to generate quality reports on fastq information. In fact, this is ordinarily the get-go thing y'all should practise once you receive a new dataset. FastQC reports provide a series of plots that allow the user to appraise the overall quality of their raw information and detect potential biases and issues.
Some plots indicate distribution of base of operations qualities along the length of reads. At to the lowest degree for illumina data, on average the quality of each base of operations tends to decrease forth the length of the read.
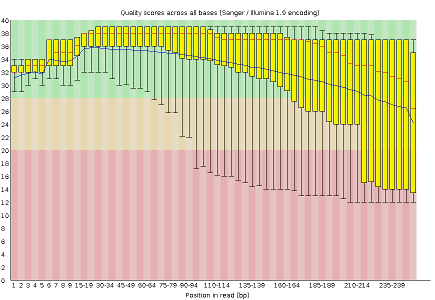
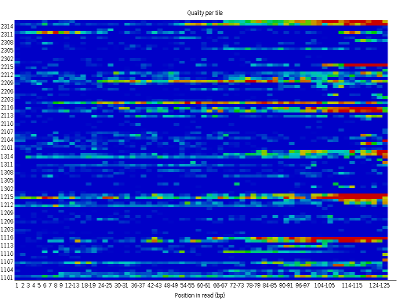
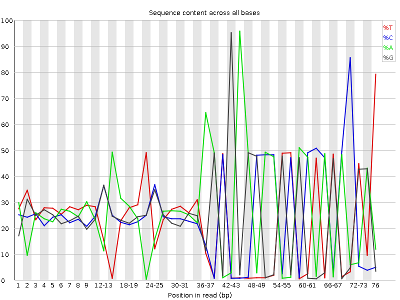
Other plots betoken biases in nucleotidic content of reads, either globally (such as %GC plots), or positionally. Global bias in nucleotidic content can be useful to search for signs of contaminants. On the other hand, positional bias are useful to detect presence of artefactual sequences in your reads such equally adaptors. Another insight you may obtain from this information are potential biases in the grooming of your library. For example, random hexamer priming is really not truly random and preferentially selects sure sequences. The currently popular transposase-based enzymatic protocol, although reasonably random, is too not completely random, and you can come across this through positional bias, particularly in the beginning of reads. The presence of adaptors is a relatively common event, and therefore specific plots exist to detect the presence of the most commonly used adaptors. Finally, the presence of repetitive sequences can too suggest contaminants, PCR artifacts, or other types of bias.
Note: Given the size of fastq files (normally in the club of Gb), they are most often compressed equally fastq.gz files. In fact, most tools (such equally FastQC) piece of work directly with fastq.gz to reduce space.
TASK: Open up a terminal. Type fastqc and press enter. The graphical interface of FastQC should appear. Open the file MiSeq_76bp.fastq.gz inside of the folder fastq_examples. Look at the dissimilar plots you obtained. Next, open the file MiSeq_250bp.fastq.gz.
QUESTION: What information is in a FastQC study?
Click Here to run across the answer
A FastQC report includes, among other things:- Basic statistics of the fastq file, including the number of reads and sequence length
- Per base of operations sequence quality, displaying the boxplot distribution of the Phred Quality (Q) per base of operations for all reads
- Per sequence quality scores displaying the histogram of the mean quality (Q value) of the bases of each read, for all reads
- Per base of operations sequence content, displaying the frequency of each nucleotide at each position of the read
- Per sequence GC content displaying the histogram of the GC frequency of each read, for all reads
- Sequence length distribution displaying the histogram of read lengths
- Sequence duplication levels displaying the histograms of the number of times reads announced with exactly the same sequence
- Overrepresented sequences (not necessarily complete reads) that appear more than frequently than randomly expected
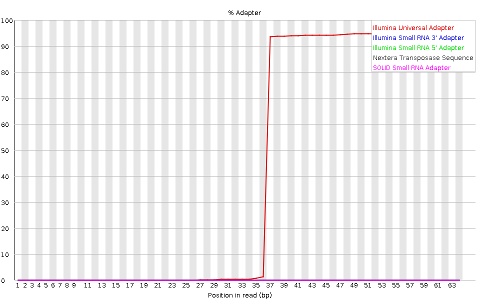
- Adapter content indicating the frequency of sequences of know sequencing adaptors forth the length of the reads
QUESTION: What are the primary differences betwixt the reports of both fastq files?
Click Hither to see the answer
The MiSeq_250bp fastq file contains 10000 reads of 250bp, while the MiSeq_76bp contains g reads of 76bp.
The MiSeq_250bp reads take a lower per base sequence quality at their end, while the reads of the MiSeq_76bp keep a good quality throughout.
The MiSeq_76bp reads contain a very noticeable nucleotide positional bias particularly after position 36. MiSeq_250bp also incorporate a bit of nucleotide positional bias, merely less and only for the first 10bp.
The MiSeq_250bp reads display an patently bimodal GC distribution, while the MiSeq_76bp reads seem closer to a single normal distribution.
Finally, MiSeq_76bp incorporate a clear presence of a known Illumina adaptor afterwards position 36 (probably the reason for the nucleotide positional bias nosotros saw before), while MiSeq_250bp contain a much smaller frequency of another Illumina adaptor towards the ends of the reads.
TASK: Insider the folder fastq_examples, you can come across fastq files from different sequencing technologies or applications. In a terminal window, go to the folder fastq_examples. Type and press enter: fastqc *.fastq.gz
Inside the folder, you lot should now see a series of HTML files with FastQC reports of each of the fastq files. Y'all can open up them with the spider web browser past clicking on them with the mouse, or by running: firefox *.html
QUESTION: Can you come across differences between the different sequencing technologies?
Click Here to see the answer
Illumina machines generate shorter reads, all with the aforementioned length. Pacbio and nanopore generate (much) longer reads, with diverse read lengths, and of a poorer quality. Illumina generates many more reads, making both technologies complementary to each other (this will become clearer when we wait at specific applications). Finally, y'all can likewise notice that, independently of the engineering science, the quality of base quality tends to subtract along the length of the read.QUESTION: What is the major divergence betwixt the two paired fastq files of the paired_example?
Click Here to see the answer
The reverse read has poorer quality bases. This is unremarkably the case, at least for illumina. This is because the reverse reads are generated later on the forward reads.Note: Assess how well you achieve the learning outcome. For this, see how well you lot responded to the different questions during the activities and likewise brand the following questions to yourself.
-
Could you run FastQC on a fastq file?
-
Can you lot broadly list types of information that a FastQC report contains?
-
Can you translate information in a FastQC written report to detect potential issues with information in a fastq file?
9.iii - Filtering and Trimming
Every bit you may take noticed before, reads tend to lose quality towards their end, where there is a college probability of erroneous bases existence called. To avoid bug in subsequent analysis, you should remove bases with higher probability of error, ordinarily by trimming poor quality bases from the end.
Task: Manually remove the bases with Q<30 from the three' end of the read you analysed earlier.
@SRR022885.ane BI:080102_SL-XAR_0001_FC201E9AAXX:vi:ane:752:593/1 CGTACCAATTATTCAACGTCGCCAGTTGCTTCATGT + IIIIIIIIII>IIIIIII@IIII.I+I>35I0I&+/ QUESTION: How is the read after this trimming operation?
Click Here to see the answer
@SRR022885.1 BI:080102_SL-XAR_0001_FC201E9AAXX:six:1:752:593/1 CGTACCAATTATTCAACGTCGCCAGTTGCTTCA + IIIIIIIIII>IIIIIII@IIII.I+I>35I0IQUESTION: Did y'all remove all lower quality bases from the read? What other strategies you can imagine to filter your reads?
Click Hither to see the reply
- No. In that location are still low-quality bases in the read (NOTE: this does not mean the base is incorrect, just that it is more likely to be incorrect).
- Instead of looking only at the last base of operations, ane tin can look at the hateful quality of the k (eg. k=4) last bases to make up one's mind if a base should be removed or not. Another alternative that is often used is to discover the longest continuous stretch of bases with a quality above a certain value.
QUESTION: Can you remove bases in the middle of reads? Why?
Click Here to meet the respond
NO! Because you would exist making artificial deletions in the sequence.Similar you have FastQC to automatically produce plots from fastq files, you lot as well accept software to filter low-quality bases from fastq files. Seqtk is a very simple tool that you can use to perform this filtering.
Job: In a terminal, go to folder fastq_examples. Type seqtk trimfq -q 0.01 MiSeq_250bp.fastq.gz > MiSeq_250bp.trimmed.fastq
QUESTION: What is this command doing? Utilize fastQC to check the new fastq file that is created by this command.
Click Here to see the answer
Seqtk removes bad quality bases from the ends of reads. In this case, it removes bases with a probability of error greater than 1% (0.01), corresponding to Q<20.Near software for the analysis of HTS data is freely bachelor to users. Nonetheless, they often require the use of the control line in a Unix-like environment (seqtk is one such example). User-friendly desktop software such as CLC or Ugene is bachelor, but given the quick step of development in this expanse, they are constantly outdated. Moreover, even with better algorithms, HTS analysis must often exist run in external servers due to the heavy computational requirements. One popular tool is Galaxy, which allows fifty-fifty non-expert users to execute many different HTS analysis programs through a simple web interface.
Task: Let'south employ Galaxy to run some bioinformatic tools. Open the web browser (eg. Firefox). Type localhost:8080 in the URL tab (where you put the web addresses). This means that y'all are accessing a galaxy example that is running on your local machine. You should come across the Milky way interface on your web browser. Click on the upload icon on the top left of the interface. Upload into Galaxy the files MiSeq_76bp.fastq.gz and MiSeq_250bp.fastq.gz. You should now seem them on your history in the right console. You lot can visualize their content by pressing the view data icon (the centre icon). After y'all have your data, you're gear up to run some tools on your data. The tools are listed on the left panel. Search for fastqc on the tool search bar on the left panel. By clicking on the tool you lot should accept in the heart the interface to run fastQC. To run fastc y'all merely need to select the fastq file and printing "Execute". Run fastqc on the fastq files you uploaded and see the result. Nonetheless in galaxy again, search for and run "seqtk trimfq" on the file MiSeq_250bp.fastq with the same parameters equally you used in the control line.
As we saw earlier, sequencing machines (namely, the illumina ones) crave that yous add specific sequences (adaptors) to your Dna so that it can exist sequenced. For many different reasons, such sequences may end up in your read, and you need to remove these artifacts from your sequences.
QUESTION: How can adaptors appear in your sequences? Have the sample MiSeq_76bp.fastq.gz every bit an example.
Click Here to see the respond
When the fragment being read is smaller than the number of bases the sequencing machine reads, so it will start reading the bases of the adaptor that is fastened to all fragments and then they can exist read past the machines. In the case of MiSeq_76bp, the fragments were all 36bp, and since 76bp were beingness read, the remaining bases belong to the illumina adaptor that was used.There are many programs to remove adaptors from your sequences, such as cutadapt. To use them y'all need to know the adaptors that were used in your library grooming (eg. Illumina TruSeq). For this, y'all need to inquire the sequencing center that generated your data.
TASK: In Milky way, employ cutadapt to remove adaptors from MiSeq_76bp.fastq.gz. In this sample, nosotros know that we used the illumina adaptor GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT, and so endeavour to remove this from the 3' end of reads and run into the touch on of the procedure using FastQC. For this, you demand to insert a new adapter in 3', and in the source, select "Enter a custom sequence" (yous don't demand to add a name, simply paste the sequence).
QUESTION: What happened? To answer, expect at the study from cutadapt, and apply FastQC on the fastq that is output by cutadapt.
Click Here to see the reply
Most no read was afflicted. This is because what you become is a readthrough, and so what is actually in the read is the contrary complement of the adaptor. Now, try the same procedure but with AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC (reverse complement of the previous). This time, nearly reads should have had the adaptor removed.Trimmomatic is a tool that performs both trimming of low-quality reads, as well as adaptor removal. Moreover, it already contains a library of commonly used adaptors, so you lot don't need to know their sequence. Like to FastQC, it is a java program, so you lot tin use it in any operating system (such equally Windows and Mac), although unlike FastQC it needs to be run only using the command line.
QUESTION: Find and select the Trimmomatic tool in Galaxy. What different operations tin can you perform with Trimmomatic that use the base quality information?
Click Here to encounter the reply
</br> You lot can perform the following operations with Trimmomatic (either isolated or in combination):- ILLUMINACLIP: Cut adapter and other illumina-specific sequences from the read
- SLIDINGWINDOW: Perform a sliding window trimming, cutting once the average quality inside the window falls below a threshold
- MINLEN: Drop the read if it is beneath a specified length
- LEADING: Cutting bases off the start of a read, if beneath a threshold quality
- TRAILING: Cutting bases off the end of a read, if below a threshold quality
- CROP: Cut the read to a specified length
- HEADCROP: Cutting the specified number of bases from the start of the read
- AVGQUAL: Drop the read if the average quality is below a specified value
- MAXINFO: Trim reads adaptively, balancing read length and fault rate to maximise the value of each read
TASK: Let'southward use Trimmomatic in Galaxy to remove low-quality bases from MiSeq_250bp.fastq.gz, as well as the remainings of illumina Nextera adaptors that are notwithstanding left in some of the reads. Now Trimmomatic should find it. Let'due south perform the default operation "SLIDINGWINDOW" of size iv and average quality 20, and then also the performance "MINLEN" using 36 every bit the minimum length. Let's also remove the adaptors. For this, select 'Yes' on 'Perform initial ILLUMINACLIP step'. And then select "Nextera (paired-end)" and leave the rest of the parameters every bit they were. Finally, click on Execute.
QUESTION: What happened? To answer, employ FastQC on the fastq output by Trimmomatic.
Click Here to meet the answer
The base quality distribution improved. Moreover, the few Nextera primers at the end of the reads also disappeared. Nonetheless, read length is now a range, and nosotros have fewer and shorter reads than before.Job: Let's employ Trimmomatic in Milky way with a paired-finish dataset. Upload the files paired_end_example_1.fastq.gz and paired_end_example_2.fastq.gz. In Trimmomatic, select 'Paired-end (two separate input file)'. Perform the same operations every bit earlier.
QUESTION: Now, you get 4 files equally output from Trimmomatic. Can yous explain what these are?
Click Hither to see the answer
You get the post-obit paired files: Trimmomatic on paired_end_example_1.fastq (R1 paired) and Trimmomatic on paired_end_example_2.fastq (R2 paired).
These incorporate the paired reads that "survived" the quality operation from both the forrad and the opposite and could, therefore, be kept as pairs. And so, you accept the cases where just one of the pairs was removed considering of low quality. In this case, it cannot be kept every bit a pair, but in a split up "isolated" file, both for the forward (Trimmomatic on paired_end_example_1.fastq (R1 unpaired)) and the reverse (Trimmomatic on paired_end_example_2.fastq (R2 unpaired)).
QUESTION: From the "isolated" reads resulting from Trimmomatic, which one has more reads? Why is that?
Click Hither to run into the answer
The forward file (Trimmomatic on paired_end_example_1.fastq (R1 unpaired)) has more reads, because the reverse reads have usually less quality, and therefore are more likely to be removed in the filtering procedure.NOTE: Plow on the green light when you're finished. Assess how well yous reach the learning upshot. For this, see how well you lot responded to the dissimilar questions during the activities and too make the following questions to yourself.
-
Could you manually remove low-quality bases from the finish of a read in the fastq format?
-
Did y'all broadly sympathise the challenges of removing bad quality bases from reads?
-
Could you use seqtk to remove low-quality bases from the end of reads in a fastq file?
-
Did you broadly understand the challenges of removing adaptors from reads?
-
Could you use cutadapt to remove adaptors from reads in a fastq file?
-
Could you utilise trimmomatic to remove bad quality bases and remove adaptors from reads in a fastq file?
-
Did yous understand the upshot of manipulating paired-finish fastq files?
Back
Back to first folio.
halvorsenwitoomards.blogspot.com
Source: https://gtpb.github.io/ELB19F/pages/L09
0 Response to "How to Split Reads Into Forward and Reverse"
Publicar un comentario